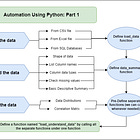
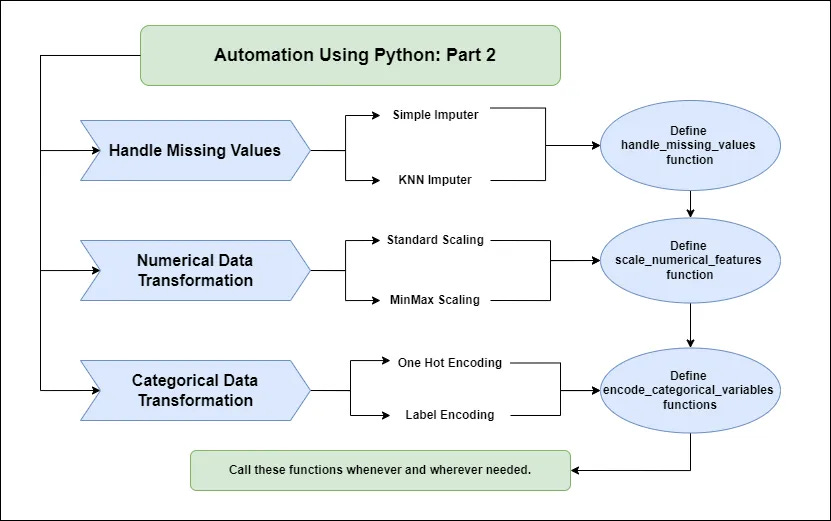
How To Automate Data Science Tasks With Python (Part 2)
In Part 2: It is about handling missing values and data transformation
This is the second article of the series “How to Automate Data Science Tasks Using Python.” If you haven't read the first part, start with this article.
What did we learn from the first article? - Tell me in the comments below. I want to know: “How different are the takeaways of each reader from the article.”
I'll tell you what I intended to educate you there:
If any task seems repeated and redundant in your project. Always define a function and automate your work.
Why we are learning this?
We are all aware that some of the tasks on our data science worklist are redundant and can be automated.
The only reason we want to automate these tasks is to save time by not having to write the same code every time.
But how can we automate? Using AI tools? No! -We will automate the tasks by defining a function and calling it as needed.
Some of the tasks we can automate include:
Data loading and understanding the data
Handling missing values and data transformation
Outlier detection and handling them
Exploratory data visualisation
Feature selection and importance
In this article, we are going to do the same thing as what we did earlier. But, here we will be creating automation functions for the second task i.e. handling the missing values and transforming the data.
Before going into the main thing, I would love it if you check out my eBooks and support me:
Also, get free data science & AI eBooks: https://codewarepam.gumroad.com/
Handling Missing Values
Following the previous article (Part 1), let us assume we have already investigated the data and are aware that there are missing values.
What should we do now?
We either drop them if the volume of missing data is low, or we impute missing values with numerical values such as mean, median, and mode.
We sometimes utilize KNN models to approximate the missing value based on the data.
This is what we usually do, right?
Imputing the missing data with mean, median or mode:
from sklearn.impute import SimpleImputer
# mean imputation
imputer = SimpleImputer(strategy = 'mean')
#median imputation
imputer = SimpleImputer(strategy = 'median')
#mode imputation
imputer = SimpleImputer(strategy = 'mode')df['column'] = imputer.fit_transform(df['column'])Imputing the missing data with KNNImputer with an estimated value:
from sklearn.impute import KNNImputer
imputer = KNNImputer(n_neighbors = 5)df[column] = imputer.fit_transform(df[column])Here, we can see that we apply multiple imputation techniques based on our needs. So, when developing a function to automate this task based on what is needed, we must include an "if-else" expression.
This is how it should be done:
from sklearn.impute import SimpleImputer, KNNImputer
# Function to handle missing values
def handle_missing_values(df, strategy='mean', columns=None, knn_neighbors=5):
if strategy in ['mean', 'median', 'mode']:
imputer = SimpleImputer(strategy=strategy)
elif strategy == 'knn':
imputer = KNNImputer(n_neighbors=knn_neighbors)
else:
raise ValueError("Unsupported imputation strategy. Choose from 'mean','median','mode', 'knn'.")
columns = df.columns if columns is None else columns
df[columns] = imputer.fit_transform(df[columns])
return dfData Transformation
What is data transformation? - It is the process of modifying data to solve a business problem and improve the performance of a model or analysis.
It is essential to note that there are two types of data forms. One is numerical, while the other is categorical data.
Numerical features:
When transforming numerical data, we normally scale it in the same range so that the model is not confused. Because the scale of different features in our numerical data might vary greatly, this can have a substantial impact on the model's performance.
Here is what we usually do for numerical data transformation:
Segregating the numerical columns first
columns = df.select_dtypes(include = ['number']).columnsPerforming standard scaling or min-max scaling according to the needs
# Standard Scaling
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()df['column'] = scaler.fit_transform(df['column'])# Min Max Scaling
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()df['column'] = scaler.fit_transform(df['column'])These are the codes we must write repeatedly for each numerical feature in our dataset.
To automate this, we can develop a function that combines all of these instructions, so that all of the numerical data can be scaled with a single call.
Something like this:
from sklearn.preprocessing import StandardScaler, MinMaxScaler
def scale_numerical_features(df, scaling_type = 'standard', columns = None):
if columns is None:
columns = df.select_dtypes(include = ['number']).columns
if scaling_type = 'standard':
scaler = StandardScaler()
elif scaling_type = 'minmax':
scaler = MinMaxScaler()
else:
raise ValueError("Unsupported scaling type. Choose 'standard' or 'minmax'.")
df[columns] = scaler.fit_transform(df[columns])
return df
Categorical features:
And, for categorical data, we should keep in mind that machine-like computers typically function well with numbers, and to improve the effectiveness of our model or analysis, we ought to present this data in some kind of numerical representation. That is where we do the encoding.
We encode in two ways: using the most common “OneHotEncoder” and with the “LabelEncoder.”
We utilize “LabelEncoder” when our categorical data is ordinal; if it is not ordinal but nominal, we use “OneHotEncoder.”
Let’s see this is how we perform encoding of any categorical data:
One Hot Encoder:
columns = df.select_dtypes(include = ['object']).columns# Using Pandas
import pandas as pd
df = pd.get_dummies(df, columns = columns, drop_first = True)# Using Scikit-learn
from sklearn.preprocessing import OneHotEncoder
encoder = OneHotEncoder(sparse_output = False)
df['column'] = encoder.fit_transform(df['column'])Label Encoder:
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
df['column'] = encoder.fit_transform(df['column'])Am I doing this correctly? Isn't the aforementioned process what we typically do?
Now, how can we automate this? To automate the entire process, we will integrate both logics into a single function called "encode_categorical_variables".
Like this:
import pandas as pd
from sklearn.preprocessing import OneHotEncoder, LableEncoder
def encode_categorical_variables(df, encoding_type = 'onehot', columns = None):
if columns is None:
columns = df.select_dtypes(include = ['object']).columns
if encoding_type == 'onehot':
df = pd.get_dummies(df, columns = columns, drop_first = True)
elif encoding_type == 'label':
label_encoders = {}
for col in columns:
le = LabelEncoder()
df[col] = le.fit_transform(df[col])
label_encoders[col] = le
else:
raise ValueError("Unsupported encoding type. Choose 'onehot' or 'label'.")
return dfWrapping it Up:
We have now created multiple separate functions for handling missing values, scaling numerical features, and encoding categorical features.
It's time to employ these functions to automate operations at any stage of the process.
We have these functions with us. Whenever and whenever missing values and data modification are required. We can just call the functions like this:
df = handle_missing_values(df, strategy = 'median')
df = scale_numerical_features(df, scaling_type = 'minmax')
df = encode_categorical_variables(df, encoding_type = 'onehot')Was this helpful? If you find this helpful, please consider:
❤ Like this article.
Subscribe to Your Data Guide for the remaining parts.
If any task seems repeated and redundant in your project.
Always define a function and automate your work
Connect: LinkedIn | Gumroad Shop | Medium | GitHub
Subscribe: Substack Newsletter | Appreciation Tip: Support